Target Identification and Evaluation in the Era of Artificial Intelligence
![]() 04/23 2026
04/23 2026
![]() 343
343
-01-
Introduction
Artificial intelligence (AI) is emerging as a transformative tool in drug discovery and development, influencing every stage from target identification to clinical trial outcome analysis. Target identification is the first step in discovering and developing new drugs, aiming to select a biological molecule that can be modulated to produce the desired therapeutic effect with adequate safety. This step is crucial for the success probability and resource requirements of subsequent steps.
Traditional target discovery remains challenging, primarily due to limited understanding of the biological complexity underlying many diseases and the technical and resource constraints in extracting necessary insights from sources such as human genomics and disease models. Out of approximately 20,000 protein-coding genes in humans, an estimated 4,500 are considered 'druggable.' However, all approved drugs to date act through only 716 distinct targets, representing a small fraction of the druggable target space and highlighting significant opportunities for future target discovery research.
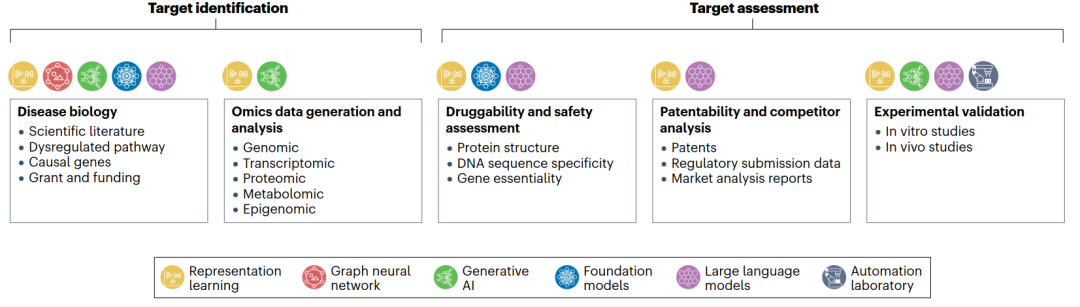
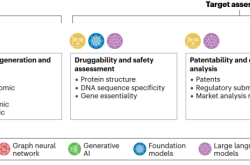
Therapeutic target identification involves selecting a disease area, followed by comprehensive mining and analysis of various existing data sources related to disease biology, and extends to generating, acquiring, and analyzing new data to compile a list of potential targets. These candidate targets must then undergo rigorous evaluation based on key criteria such as druggability, safety, patentability, competitive analysis, and experimental validation. The efficiency and effectiveness of many steps can be enhanced by suitable AI tools capable of integrating heterogeneous large-scale datasets, constructing a unified representation of disease biology, and uncovering underlying mechanisms through machine learning models. Furthermore, AI can predict key characteristics related to target accessibility and risk, and guide experimental design through automated laboratories, enabling iterative optimization.
-02-
I. Key Considerations for Target Identification
1. Therapeutic Hypothesis
Identifying a therapeutic target and its mechanism of modulation expected to influence disease biology, translate into meaningful therapeutic effects for patients, and possess adequate safety is a complex and multifaceted challenge. Historically, studies in animal disease models and/or clinical observations have often provided critical information for formulating therapeutic hypotheses around specific biological targets. With advances in genomic technologies since the 1980s, knowledge from human genetics and/or studies involving genetic perturbations in cellular and animal models has become an increasingly important contributor to target identification efforts.
Over the past 15 years, there has been a growing recognition of the limited predictive value of animal models for many diseases and the importance of human-derived evidence supporting therapeutic target hypotheses. Building on this, a notable trend is the application of causal biological research, employing causal inference techniques in large human datasets. Researchers integrate genome-wide association studies (GWAS) with quantitative trait locus analysis and use methods such as Mendelian randomization to examine the effects of gene or protein expression levels on disease, providing valuable evidence for potential drug targets. Another strategy in causal biological research is leveraging perturbation experimental data from human cell lines to infer mechanistic relationships and nominate therapeutic targets.
2. Druggability and Safety
The term target druggability is used to describe the potential to identify a drug candidate capable of modulating the function of a target in a manner hypothesized to lead to desired outcomes for patients with a specific disease. For small-molecule therapies, druggability is closely related to ligandability, which refers to the presence of well-defined, accessible pockets on the target's surface with which drug-like small molecules can bind. Understanding the target's three-dimensional structure and ligandability helps prioritize potential targets and guides structure-based drug design.
Today, a growing number of therapeutic modalities beyond small molecules have been clinically validated, including antibody-based therapies, oligonucleotide-based therapies, and gene and cell therapies. These modalities expand the landscape of disease-relevant targets potentially amenable to drugging. Emerging AI tools such as AlphaFold3 can be used to facilitate druggability assessments, providing accurate structural predictions for protein-ligand, antibody-antigen, and oligonucleotide-protein complexes.
While druggability is a measure of a target's susceptibility to clinical perturbation by a therapeutic agent, safety is inherently related to the specificity of this interaction. Assessing the risk of such effects during the target identification phase relies on predictive toxicology and analysis of target-related biological pathways to anticipate potential adverse effects. However, most side effects occur through off-target interactions. Although comprehensive prediction of off-target effects remains challenging, the potential risk of certain off-target interactions can be predicted during the target identification/evaluation phase based on structural similarities between the intended target and off-targets.
3. Novelty, Confidence, and Commercial Accessibility
The strategic selection of targets often involves a delicate trade-off between novelty and confidence in the target's role in disease. High-confidence targets are supported by more scientific evidence, offering a clearer path to clinical translation and reducing the risk in drug development. However, novel targets provide opportunities for breakthrough therapies, particularly for diseases with unmet medical needs.
The interplay between novelty and confidence relates to the commercial accessibility of targets. Novel targets have the potential to revolutionize disease treatment and open up new markets, while established targets offer more predictable and safer investments but face greater competition from other drugs. This balance, often referred to as 'first-in-class' versus 'best-in-class,' influences market dynamics, competitiveness, intellectual property strength, overall investment, and potential returns.
4. Combination Value
The potential of a target to provide the basis for a drug administered as part of a combination therapy is often increasingly important, as the synergistic or additive effects of drug combinations have been shown to reduce the likelihood of resistance, enhance therapeutic efficacy, and broaden therapeutic applications.
-03-
II. Application of AI in Therapeutic Target Identification
1. Data for Target Identification
The cornerstone of AI's application in identifying therapeutic targets is its ability to process and analyze a wide range of complex, multimodal data. Publicly available data sources include omics data, biological maps, clinical and phenotypic data, textual information, and intellectual property and industry information.
Omics Data: Includes genetic, transcriptomic, proteomic, metabolomic, epigenomic, and microbiome data, providing comprehensive biological information that enables a systematic view of the molecular aspects of diseases. Machine learning models facilitate the integration of diverse multi-omics data to construct a more holistic molecular map of diseases.
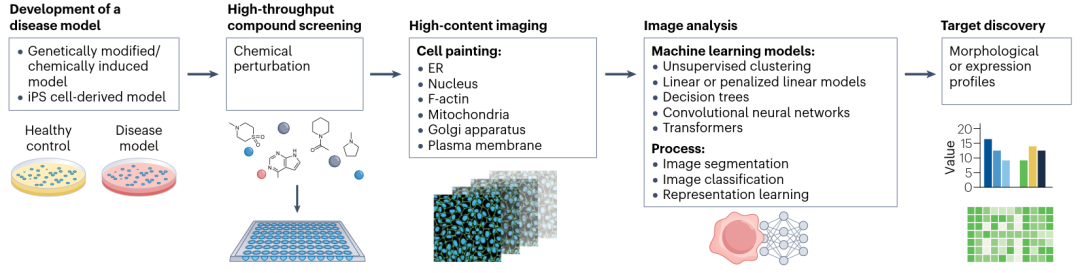
Cellular Imaging Data: AI-based biological image analysis can inform target identification, drug screening, and optimization of potential therapies. For example, the Cell Painting assay, combined with machine learning algorithms, has been used to identify therapeutic targets for intestinal fibrosis.
Biological Knowledge Graphs: Biological knowledge graphs organize and represent complex relationships and interactions among biological entities in a graph structure. Various types of knowledge graphs have become popular data sources for AI-based target research.
Clinical and Phenotypic Data: When analyzed in conjunction with AI, these data can reveal insights into disease mechanisms, patient stratification, and potential therapeutic pathways.
Textual Information: AI systems can parse and analyze vast amounts of scientific literature to extract relevant information, generating new disease hypotheses, identifying therapeutic targets, and formulating treatment strategies.
Integrating Heterogeneous Data Types: To maximize the value of heterogeneous data sources for target identification and evaluation, effective integration strategies are crucial. One approach is to construct heterogeneous knowledge graphs; another strategy is to use database technologies to orchestrate heterogeneous data sources within a single platform or data warehouse.
2. AI Models for Target Identification
Machine learning demonstrates significant potential in analyzing vast biomedical data to discover previously unknown disease-related targets. Machine learning frameworks for target identification and evaluation include supervised learning, unsupervised learning, and semi-supervised learning. Additionally, emerging AI technologies such as representation learning, graph neural networks, generative AI, foundational models, and large language models are also applied in target discovery.
Supervised Learning: Supervised learning uses labeled data to train models, enabling them to make predictions or classifications on unseen datasets. This framework is widely used for predicting drug-target interactions. Supervised learning methods are also employed to predict targets for specific diseases by incorporating target features.
Unsupervised Learning: Unsupervised learning analyzes unlabeled data to identify hidden relationships and structures, often using techniques such as clustering and dimensionality reduction. Another form is self-supervised learning, which leverages intrinsic signals within unlabeled data to generate supervisory information, enabling the model to learn meaningful patterns without external annotations.
Semi-Supervised Learning: Semi-supervised approaches combine a small amount of labeled data with a large amount of unlabeled data, often helping to improve the learning process on large unlabeled datasets when labeled data is scarce or costly to acquire.
Representation Learning: Representation learning underpins advanced AI technologies such as graph neural networks, generative AI, foundational models, and large language models. It encodes different biological data entities into latent embedding vectors that capture intrinsic and context-dependent biological properties, facilitating a wide range of downstream tasks including target identification.
Graph Neural Networks: Graph neural networks enable AI systems to leverage biological map data for target discovery.
Generative AI: Generative AI models, often built on deep neural network architectures, aim to generate new data and have been widely applied in areas such as text and image generation. In the context of target identification, generative AI can be used to generate synthetic datasets, facilitating downstream analysis.
Foundational Models: Foundational models have achieved remarkable success in natural language processing and computer vision by pre-training on vast, diverse datasets. After fine-tuning, these models are highly versatile and often outperform task-specific models. With the continuous growth of biomedical datasets, foundational models are expected to play an increasingly significant role in future target identification research.
Large Language Models: Large language models have spurred the emergence of AI agent frameworks for therapeutic target identification and evaluation.
3. AI Model Validation
The validation of AI models for therapeutic target identification is a critical step to ensure their reliability and applicability. Key validation methods include retrospective validation, experimental validation, and prospective validation.
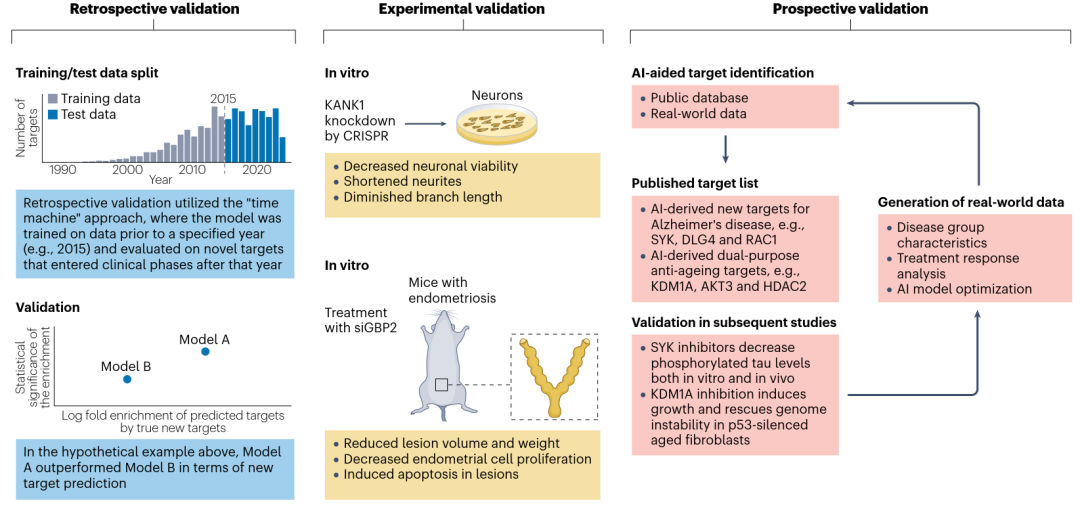
Retrospective Validation: Backtesting is a retrospective validation technique used to assess an AI model's ability to accurately identify known therapeutic targets using historical data.
Experimental Validation: Experimental validation of AI-predicted target functions and their potential interactions with therapeutic agents involves rigorous in vitro assays. In vivo animal studies can further elucidate the target's role in disease mechanisms and the potential impact of intervention strategies.
Prospective Validation: While experimental validation aims to immediately assess the biological relevance and therapeutic potential of AI-identified drug targets through experiments, prospective validation is essential to ensure that AI models can accurately predict outcomes in real-world settings. The ultimate validation of therapeutic targets lies in successful human clinical trials.
-04-III. Application of AI in Target Evaluation
1. Target Druggability Assessment
For targets selected for drug development, assessing their structural and functional characteristics, binding sites, and the feasibility of developing small molecules or biologics capable of interacting with the target in a specific, effective, and safe manner is crucial. In this regard, AI tools for protein structure prediction are highly valuable. The introduction of AlphaFold has revolutionized structural biology. Although AlphaFold has shown significant value in multiple applications, its utility in structure-based drug discovery remains uncertain.
AI can also facilitate druggability assessments by identifying cryptic pockets in proteins. These pockets, absent in ligand-free experimental structures, may form during conformational changes and have the potential to serve as binding sites for proteins previously considered undruggable.
2. Target Novelty and Commercial Accessibility
A recent study reported an AI-driven approach to evaluate cancer therapy targets through a dual assessment of novelty and commercial potential. Target novelty is quantitatively determined through a comprehensive analysis of drug development status, therapy area-specific literature, and scientific documents. Commercial accessibility is assessed using a dedicated large language model that integrates multiple parameters.
-05-IV. Examples of AI-Supported Target Identification
New targets can ultimately only be validated through successful clinical trials, which provide the basis for regulatory approval. Currently, no drugs originating from AI-driven target identification have reached this stage, with only a limited number of targets identified or candidates supported by AI tools for their therapeutic potential having entered clinical trials.
1. TNIK as a Target for Idiopathic Pulmonary Fibrosis
A recent study by Insilico Medicine demonstrated the application of end-to-end generative AI in drug discovery to identify a novel target and potential drug candidate for the treatment of idiopathic pulmonary fibrosis (IPF). Utilizing multi-omics datasets from IPF and healthy lung tissues, their AI platform ranked TNIK as a top target. Although TNIK's indirect function had been previously studied, it had not been investigated as a therapeutic target for IPF, making it a novel target selected by the platform.
2. APLNR as an Anti-Aging Target
AI-driven analysis of human data helps de-risk the clinical translation of promising drug targets. A drug discovery platform developed by BioAge incorporates longitudinal multimodal human aging datasets and applies computational tools to identify molecular signatures associated with longevity and delayed disease onset.
3. PIKfyve as a Target for Amyotrophic Lateral Sclerosis
Amyotrophic lateral sclerosis (ALS) is a rare neuromuscular disease. The therapeutic potential of PIKfyve inhibition in multiple ALS types is further supported by ConVERGE, an AI-driven drug development platform focusing on data derived from humans.
4. DRD2 as an Anti-Cancer Target
While the identification of targets such as TNIK, APLNR, and PIKfyve illustrates the target-based drug discovery paradigm, another approach involves target deconvolution following phenotypic screening. Traditionally, target deconvolution has relied on laboratory experiments, but this process is increasingly supported by AI. The identification of DRD2 as an anti-cancer drug target serves as a notable example.
-06-V. Challenges and Future Directions
The translation of computational predictions into clinically meaningful outcomes remains constrained by significant technical and operational challenges.
1. Data Quality and Availability
The application of AI in target identification heavily relies on large-scale, high-quality data for training. Despite the rapid accumulation of omics data in recent decades, its application in model training faces significant challenges. Additional limitations include the scarcity of omics data for rare diseases, racial and ethnic minorities, and socially vulnerable groups, which may result in poor model generalization. For scientific literature data, the reproducibility of published research findings poses a major challenge.
2. Integration of Multimodal Data Analysis
A promising strategy to overcome the limitations of unimodal data is the integration of multimodal datasets. Growing evidence indicates that multimodal approaches consistently outperform unimodal methods in tasks related to target discovery.
3. Explainable AI Models
Machine learning techniques, particularly deep learning, often do not provide insights into how internal computational processes lead to specific outputs, hindering the troubleshooting process and making it difficult to identify and correct errors. This opacity impedes domain experts' evaluation of these models, affecting their acceptance. As a solution to the “black box” problem, explainable models elucidate the rationale behind model predictions.
4. Standardized Metrics and Benchmarking Frameworks
The evaluation of AI models for therapeutic target identification requires standardized validation metrics and benchmarks to ensure scientific rigor and facilitate fair comparisons among methods. Standard metrics are commonly used to assess drug-target interaction prediction models. Although these metrics are effective for quantifying prediction accuracy, they often overlook the biological relevance and clinical feasibility essential for disease-specific therapeutic targets. Emerging benchmarking systems aim to overcome these limitations by integrating multimodal data and emphasizing clinical utility.
5. Synthetic Data and Digital Twins
Synthetic data refers to artificially generated data that simulates real-world biological patterns and characteristics, which can be generated by AI algorithms to mimic different biological scenarios. Synthetic data can also be used to create digital twins, which are virtual models of individual entities (e.g., patients, organs, or cells).
6. AI-Driven Closed-Loop Experimental Platforms
AI-driven closed-loop experimental platforms represent an emerging paradigm in which AI models nominate therapeutic targets, automated laboratories test target modulation, results flow into AI analysis tools, and analytical outputs are fed back into target identification models to optimize target prioritization. The closed-loop design is achieved through end-to-end robotic handling of compounds, biological samples, imaging, and multi-omics data, combined with AI tools.
-07-Conclusion
The integration of AI is enabling a more systematic and data-driven approach to therapeutic target identification and evaluation. The ability of AI to analyze multimodal data spanning high-dimensional omics, phenotypic imaging, and large-scale knowledge graphs addresses long-standing barriers related to biological complexity and translational failure. Advances in machine learning frameworks, neural architectures, generative AI, and foundational models further empower researchers to discover novel therapeutic targets and assess their druggability. This progress is reflected in several AI-identified targets that have entered clinical-stage development.
However, realizing the full potential of these technologies requires addressing persistent challenges. Overcoming data scarcity and bias, improving model interpretability, and establishing rigorous benchmarking standards remain crucial for integrating the predictions of AI tools into decision-making processes. The future of target discovery lies in the fusion of computational and experimental workflows through AI-driven closed-loop platforms, accelerating the translation of biological insights into effective, clinically viable therapies.
References: Target identification and assessment in the era of AI. Nat Rev Drug Discov. 2026 Apr 20.